View all of our scientific posters presented at the 2022 Association For Cancer Research (AACR) Annual Meeting. Or you can select a application area of your interest below.
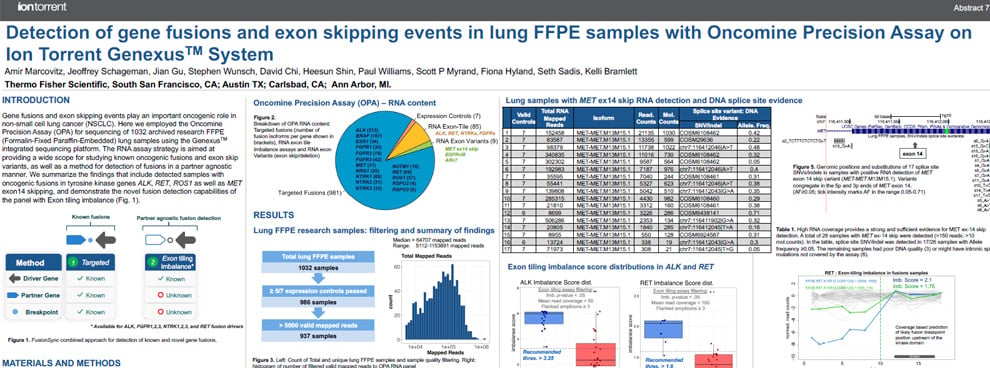
This poster describes the study of studying known oncogenic fusions and exon skip variants, as well as a method for detection of novel fusion combinations and detection of fusions in a partner agnostic manner.
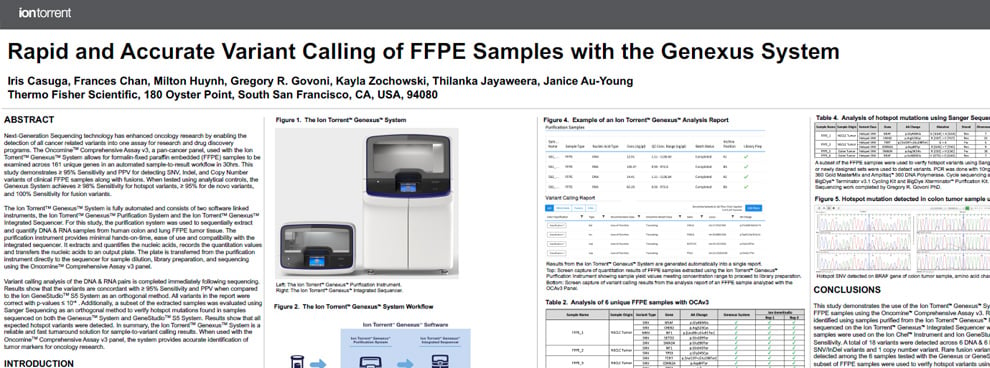

This poster demonstrates that the Genexus System is a reliable and fast turnaround solution for sample-to-variant calling results. When used with the Oncomine™ Comprehensive Assay v3 panel, the system provides accurate identification of tumor markers for oncology research.
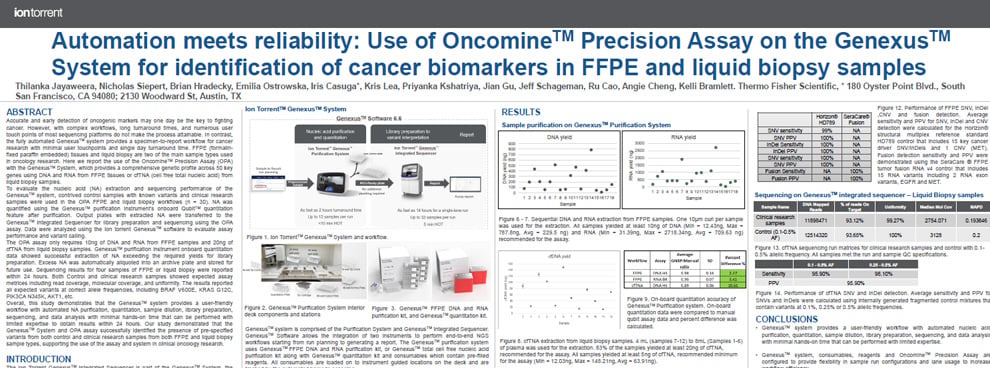

This poster demonstrates that the Genexus™ system provides a user-friendly workflow with automated NA purification, quantitation, sample dilution, library preparation, sequencing, and data analysis with minimal hands-on time that can be performed with limited expertise to obtain results within 24 hours.
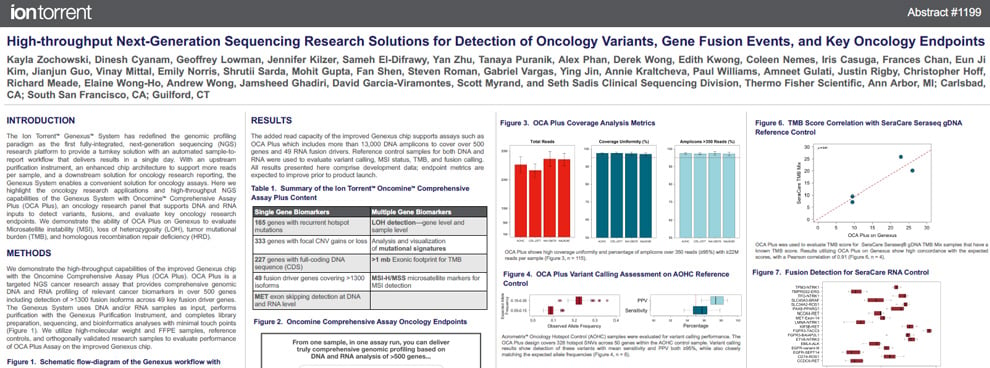
This poster contains information that demonstrates the ability to evaluate tumor mutational burden (TMB), microsatellite instability (MSI), loss of heterozygosity (LOH), and homologous recombination repair deficiency (HRD) using the Oncomine Comprehensive Assay Plus run on the Genexus System.
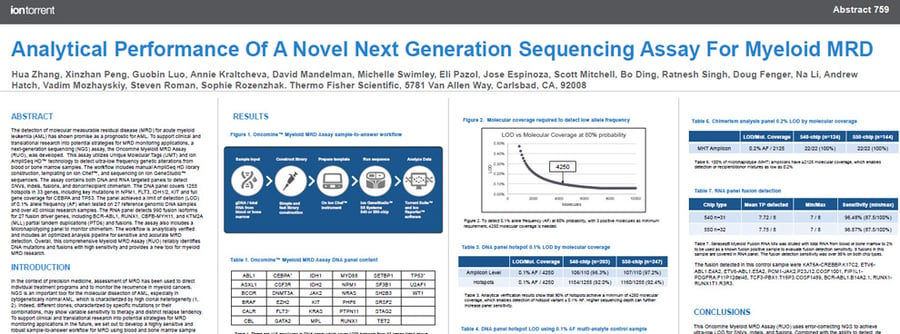
This poster characterizes a comprehensive Myeloid MRD (RUO) assay that reliably identifies DNA mutations and fusions with high sensitivity and provides a new tool for myeloid MRD research.
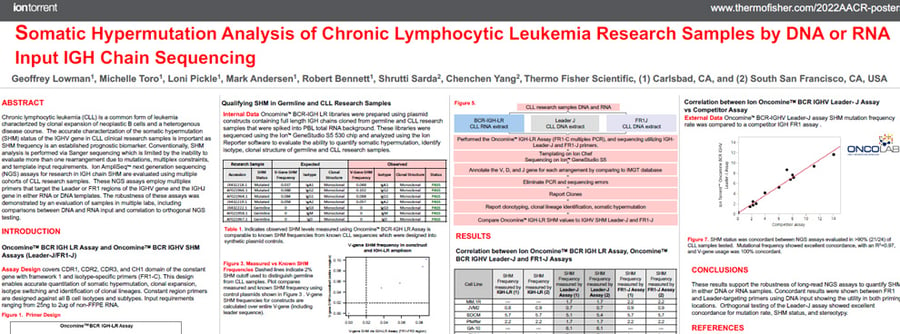
This poster describes use of Ion AmpliSeq™ next generation sequencing (NGS) assays for research in IGH chain Somatic Hyper Mutation are evaluated using multiple cohorts of CLL research samples.
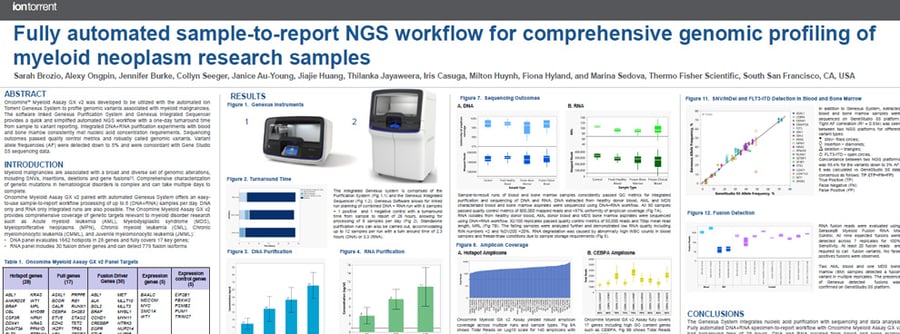
This poster describes development of a fully automated NGS Myeloid Assay that offers an easy to use sample-to-report workflow and the capability for processing up to 8 samples per day.
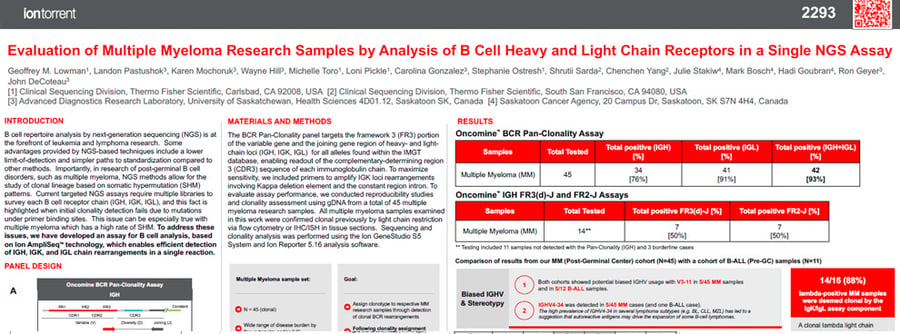

This poster describes results of B cell repertoire analysis in multiple myeloma samples by next-generation sequencing (NGS) through use of the Oncomine BCR Pan-Clonality Assay.
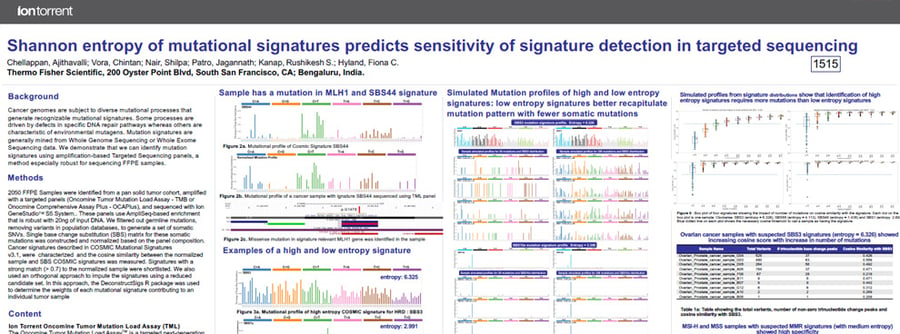
This poster describes the complexity of the COSMIC mutational signatures and their impact while introducing Shannon entropy as a metric to measure complexity of trinucleotide distributions.
For Research Use only. Not for use in diagnostic procedures.
We've detected your location to be Japan.
Sorry, you cannot access this website. The content on www.oncomine.com is only intended for healthcare professionals. Formore information on our research solutions, please visit ThermoFisher.com
このウェブサイトは、日本国内の医療関係者の方への情報提供を目的としており、一般の方に対する情報提供を目的としたものではないことをご了承ください。研究用製品の情報はThermoFisher.comよりご覧ください。