Lymphoproliferative disorders such as acute lymphoblastic leukemia (ALL), lymphomas, and multiple myeloma comprise the majority of new blood cancer cases diagnosed each year.
These diseases affect cells of the adaptive immune system, including T-cells, B-cells, and plasma cells. Molecular analysis of these neoplasms can:
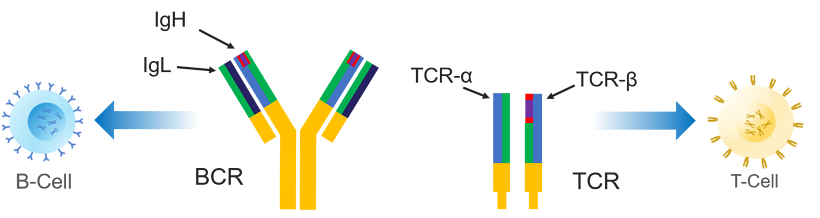
Clonality testing is an important molecular testing method that characterizes B-cell receptors (BCR, aka immunoglobulins) and T-cell receptors (TR or TCR), which serve as unique identifiers of the individual B-cell and T-cell clones in our immune system.
What does clonality testing tell you?
In a healthy blood sample, there are many different B-cell and T-cell clones, resulting in a diverse immune repertoire or a ‘polyclonal’ population. This diversity is necessary, as the immune system must be capable of responding to a wide array of antigens that are continually encountered.
The BCR and TCR cell surface receptors, whose role is to bind to foreign antigens, must be diverse in order to selectively respond. Occasionally, a single clone will expand and become dominant resulting in a monoclonal population. This expansion may be an indication of a malignant clone that has multiplied uncontrollably.
To study the diversity, or lack there-of, of the immune system, sequencing of all the cell types (BCR immunoglobulins and TCRs) is necessary.
NGS for studying the immune repertoire
Immune repertoire plays a significant role in both hemato-oncology and immune-oncology studies1,2. A tremendous amount of data can be gathered through next-generation sequencing (NGS) of genes that code for the BCR and TCR chains.
Starting with DNA or RNA from blood, bone marrow or other sample types, NGS can help establish the degree of clonal diversity, level of somatic hypermutation, detect low-frequency residual clones after drug exposure, and more.
Analysis of sequencing data has long been viewed as requiring a bioinformatics specialist. However, advances in analysis software tools have focused on providing user friendly interfaces and output tables and graphs.
References:
1. Aran, A., Garrigós, L., Curigliano, G., Cortés, J., & Martí, M. (2022). Evaluation of the TCR Repertoire as a Predictive and Prognostic Biomarker in Cancer: Diversity or Clonality?. Cancers, 14(7), 1771. https://doi.org/10.3390/cancers14071771
2. van Bladel, D., van der Last-Kempkes, J., Scheijen, B., Groenen, P., & EuroClonality Consortium (2022). Next-Generation Sequencing-Based Clonality Detection of Immunoglobulin Gene Rearrangements in B-Cell Lymphoma. Methods in molecular biology (Clifton, N.J.), 2453, 7–42. https://doi.org/10.1007/978-1-0716-2115-8_2
Precision medicine is rapidly changing our understanding of cancer research and treatment decisions. These breakthrough, personalized treatments hold promise even for patients with historically hard-to-treat diseases, like lung or breast cancer. But
In his recent talk at OncomineWorld 2023, Dr. Bekim Sadikovic presented a strong, evidence-based argument for frontline next-generation sequencing (NGS) in myeloid malignancy testing.
Bekim Sadikovic PhD, DABMGG, FACMGProfessor, Research Chair, and...
In a recent Association for Molecular Pathology (AMP) workshop, Dr. Cecilia Yeung shared how her lab is implementing rapid next-generation sequencing (NGS) to help advance hemato-oncology research and expand access to underserved populations.
...